Application of an advanced human experimental model of the liver tumor microenvironment (TME) to identify targetable metastatic phenotypes and test potential therapeutics for metastatic breast cancer and melanoma.
We are integrating the combined use of quantitative systems pharmacology (QSP) and biomimetic human liver microphysiology systems (MPS) to recapitulate the human liver acinus as an advanced human experimental model for defining targetable metastatic phenotypes and for testing therapeutics for metastatic breast cancer and both metastatic cutaneous and uveal melanoma in the liver tumor microenvironment (TME).
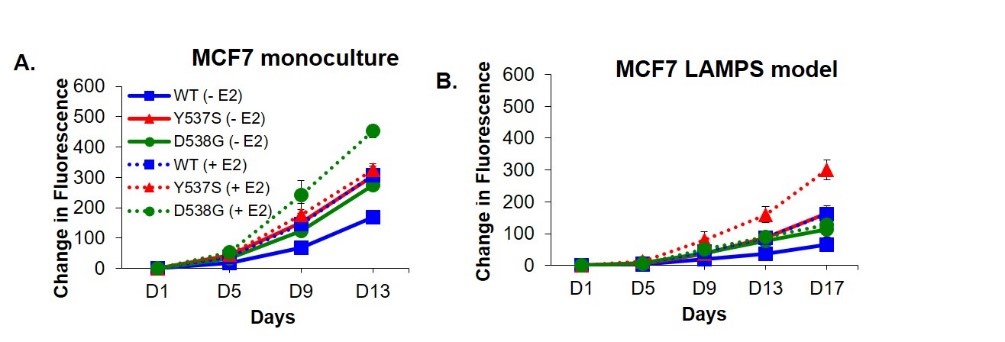
Our collaborations in the breast cancer program include Adrian Lee, PhD, Director of the Institute for Precision Medicine, and an internal advisory board member for the UPDDI, Chakra Chennubhotla, PhD, Department of Computational and Systems Biology and PredxBioTM COO, as well as Andy Stern, PhD, Bert Gough, PhD, Mark Miedel, PhD, all from the Department of Computational and Systems Biology and the UPDDI. In addition, a spin-off company, PredxBioTM, a computational and systems pathology company, was formed to commercialize the intellectual property based on spatial analytics, explainable artificial intelligence (xAI) and systems biology applied to primary tumors to generate prognostic and diagnostic tests, as well as to guide personalized therapeutic strategies.
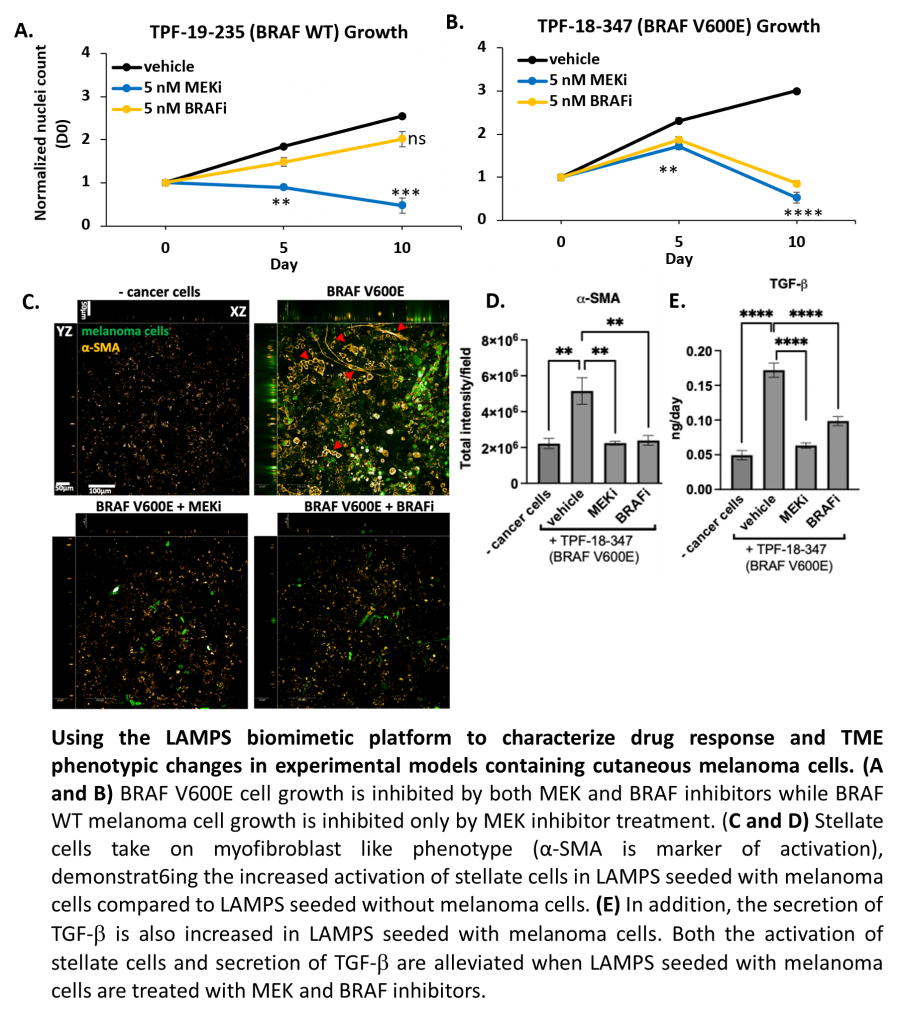
The goal of the metastatic melanoma program is to develop therapeutics for metastatic melanoma using QSP. We take advantage of the strengths in melanoma cancer within the UPMC Hillman Cancer Center with Diwakar Davar, MD, MS as clinical director for the Melanoma and Skin Cancer Program and co-Directors John Kirkwood, MD, and Hassane Zarour, MD, a member of the UPDDI internal advisory board. We also harness the strengths in cancer, biomedical engineering and organoid engineering at Vanderbilt with John Wikswo, PhD, Biomedical Engineering, Molecular Physiology & Biophysics and Physics and the University of Wisconsin with William Murphy, PhD, Department of Biomedical Engineering and Pharmacology and Co-Director of the Stem Cell and Regenerative Medicine Center. The UPDDI, Vanderbilt and Wisconsin have an NIH UO1 grant to apply the integrated technologies to develop therapeutics. Key faculty also include Alex Soto-Gutierrez, MD, PhD, Department of Pathology and the McGowan Institute of Regenerative Medicine, as well as Albert Gough, PhD, Larry Vernetti, PhD, Mark Miedel, PhD all from the Department of Computational and Systems Biology and the UPDDI.
Click here and include “Metastatic Cancer” in the Additional Gift Instructions to support development of an advanced tumor microenvironment model metastatic cancer target identification and therapy development.
